Ana Catalán - Firefly genomics and transcriptomics
The genetic basis of sexual dimorphism in fireflies
One of the most extreme differences within species are sexually dimorphic traits, many of which can be linked to sex-biased gene expression. However, the genomic basis and evolutionary processes explaining sex-biased gene expression divergence are not well understood. We use fireflies as a novel system to study sexual dimorphism and the role of transcriptome evolution for the maintenance of sexually dimorphic traits.
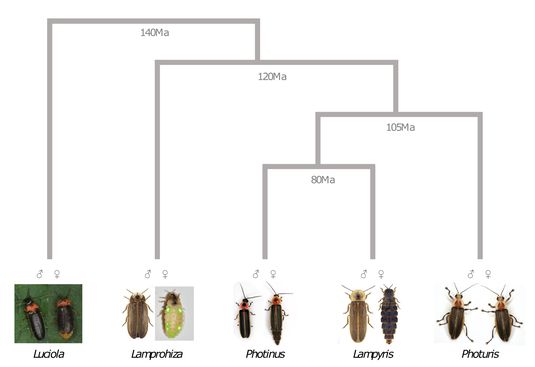
Firefly phylogenetic tree. Female neoteny has evolved independently many times, as in Lampyris and Lamprohiza.
Demography and adaptation in European fireflies
Understanding the demographic history of species is a key component to understand species colonization patterns and the adaptational events enabling colonization to novel environments. Using various population genetic approaches, our main aim is to find the genetic regions that contribute to the process of adaptation using firefly species that have a wide geographical distribution and therefore were able to adapt to different niche types.
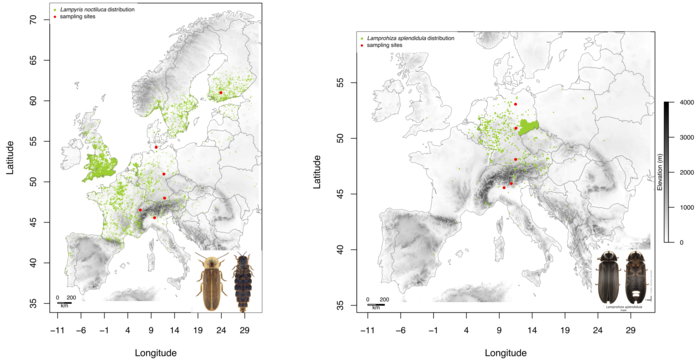
Lampyris noctiluca and Lamprohiza splendidula have an ample distribution in Europe
Firefly biodiversity in the Neotropics
Tropical regions harbor the highest biodiversity around the globe and yet, several of these regions are strongly understudied. Particularly, the study of firefly biodiversity in the Neotropics has been heavily neglected, leaving in the dark their identity and phylogenetic relationships. We are working in collaboration with del Valle University of Guatemala to uncover the biodiversity of fireflies in a highly endemic region. We are using morphological and genetic approaches for species discovery and identification.

Different Neotropical Photinus firefly species, many of which taxonomic and phylogenetic identification awaits.
Techniques
To study the evolution of genomes and transcriptomes in fireflies we are generating high quality genomes and transcriptomes using various technologies: PacBio (HiFi, Iso-Seq), Nanopore, Hi-C and Illumina short read sequencing.
Publications
See google scholar: https://scholar.google.de/citations?user=SosUN-8AAAAJ&hl=en
Contact information
email: catalan@bio.lmu.de
office: room C01.005
personal website: www.anacatalan-evolution.com

